...
At the moment of writing this page, during the 2nd meeting of the Technical Work Group of EMODnet in the Sea Aquarium of Genova in July 2017, discussions about improvements on the services are the main topic. Until now the different EMODnet lots have carried out a great job to collect and harmonize all kinds of data sets. Now it is the time to invest in harmonizing all portals so end users can find data a bit better.
At this technical meeting Deltares gave a presentation where the a specific use case presented got much attention. The Use Case was "I want to visualise the relation between the sea floor depth and the occurrence (with depth) of a specific animal (example is for Amphiura filiformis)". Some constraints were formulated, these are:
- use the metadata delivered by the central portal (http://emodnet.eu --> product catalogue)
- only by using OGC services
- only by using free searches on species names
A step wise approach was followed presented to work out the use case. Steps The steps followed are:
- go to the catalogue of EMODnet to find the data sets, these are:
- bathymetry
- biotic observations (without some specific knowledge it is hard to find data set(s) where abundance information of Amphiura filiformis can be found)
- Go to the specific sub portals via the links presented above and find out which services are available, these are:
- for bathymetry the following services are made available
WCS gives you the real data. The example will be worked out for this service. Find out more about WCS at the page WCS primer. - for biotic observations the challenge is to find abundance information on the VLIZ geoserver
Geoserver can advertise WMS, WFS and WCS information, in case of species distribution in relation to depth the real observations are the way to go. Read more about WFS at the WFS primer page. Checking the complete geoserver and filtering on Eurobis it reveals the best data set to be used is Aggregated points (stations) of EurOBIS occurences.
- for bathymetry the following services are made available
At this point insight is gained in the available sources and possible data sets that can be used. Close investigation reveals however that the links offered for Bathymetry are not really usable. Reverse engineering via the download area of interest button on the EMODnet bathymetry portal reveals that the data set with the real data is not advertised. This is shown via the get capabilities request of the WCS services provided.
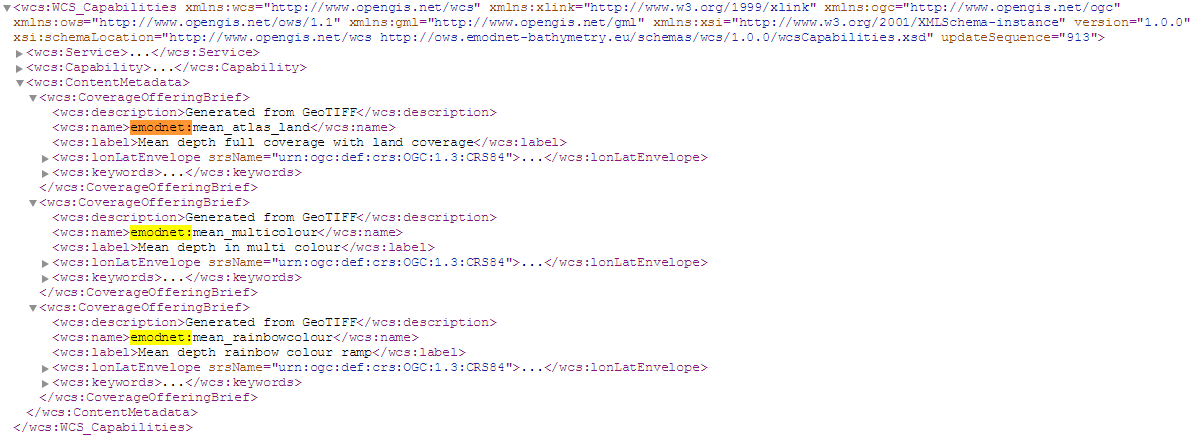
Code Block language xml title WCS GetCapabilities http://ows.emodnet-bathymetry.eu/wcs?service=WCS&request=GetCapabilities&version=1.0.0
this gives:
The developer tools of the browser used on the bathymetry data portal gives information that a different layer is used, the coverage emodnet:mean is not mentioned in the getCapabilities but is used as layer to download.
- Now a list of exact data sources is know, these are:
- for bathymetry (http://ows.emodnet-bathymetry.eu/wcs? coverage emodnet:mean)
- for biota (http://geo.vliz.be/geoserver/Eurobis/ows? layername=Eurobis:eurobis_points)
On the primer pages present code snippets for python. These code snippets are adjusted to the particular example presented here.
Code Block language py title WCS code snippet # import packages import os from owslib.wcs import WebCoverageService from osgeo import gdal # define the connection url = 'http://ows.emodnet-bathymetry.eu/wcs?' wcs = WebCoverageService(url,version='1.0.0',timeout=320) wcs.identification.type wcs.identification.title # define variables clipfile = r'..\temp.tif' requestbbox = (2.097,52.715,4.277,53.935) layer = 'emodnet:mean_atlas_land' # get the data bathyml = 'emodnet:mean' sed = wcs[layer] #this is necessary to get essential metadata from the layers advertised sed.keywords sed.grid.highlimits sed.boundingboxes cx, cy = map(int,sed.grid.highlimits) bbox = sed.boundingboxes[0]['bbox'] lx,ly,hx,hy = map(float,bbox) resx, resy = (hx-lx)/cx, (hy-ly)/cy width = cx/1000 height = cy/1000 gc = wcs.getCoverage(identifier=bathyml, bbox=requestbbox, coverage=sed, format='GeoTIFF', crs=sed.boundingboxes[0]['nativeSrs'], width=width, height=height) fn = clipfile f = open(fn,'wb') f.write(gc.read()) f.close() filetiff = gdal.Open(clipfile) theImage = filetiff.GetRasterBand(1).ReadAsArray() os.unlink(clipfile)
for querying the WFS following code is used
Code Block language py title WFS querying # import the packages import pandas import sys import os from owslib.fes import * from owslib.etree import etree from owslib.wfs import WebFeatureService # define the input variables requestbbox = (2.097,52.715,4.277,53.935) fn = r'..\afile' wfs = 'http://geo.vliz.be/geoserver/Eurobis/ows?' layer = 'Eurobis:eurobis_points' afilter = PropertyIsEqualTo(propertyname='Eurobis:ScientificName', literal='Amphiura filiformis') # define the link to the service and carry out the request wfs11 = WebFeatureService(url=wfs, version='1.1.0',timeout=320) response = wfs11.getfeature(typename=layer, bbox=bbx,srsname='urn:x-ogc:def:crs:EPSG:4326',outputFormat='shape-zip') if os.path.isfile(fn): os.unlink(fn) out = open(fn, 'wb') out.write(response.read()) out.close() # define pandas dataframe for the output out = open(fn, 'wb') out.write(response.read()) out.close() df = pandas.read_csv(fn,header=0,sep=',') os.unlink(fn)Now the result (if there is data in the boundingbox, there is no testing involved, thus the result can be empty, this is not tested) is there, in 2 objects (Pandas DataFrame with observation information and a Numpy array with bathymetry). This can be plotted in a 3D Scatter plot using the Matplotlib. See next code.
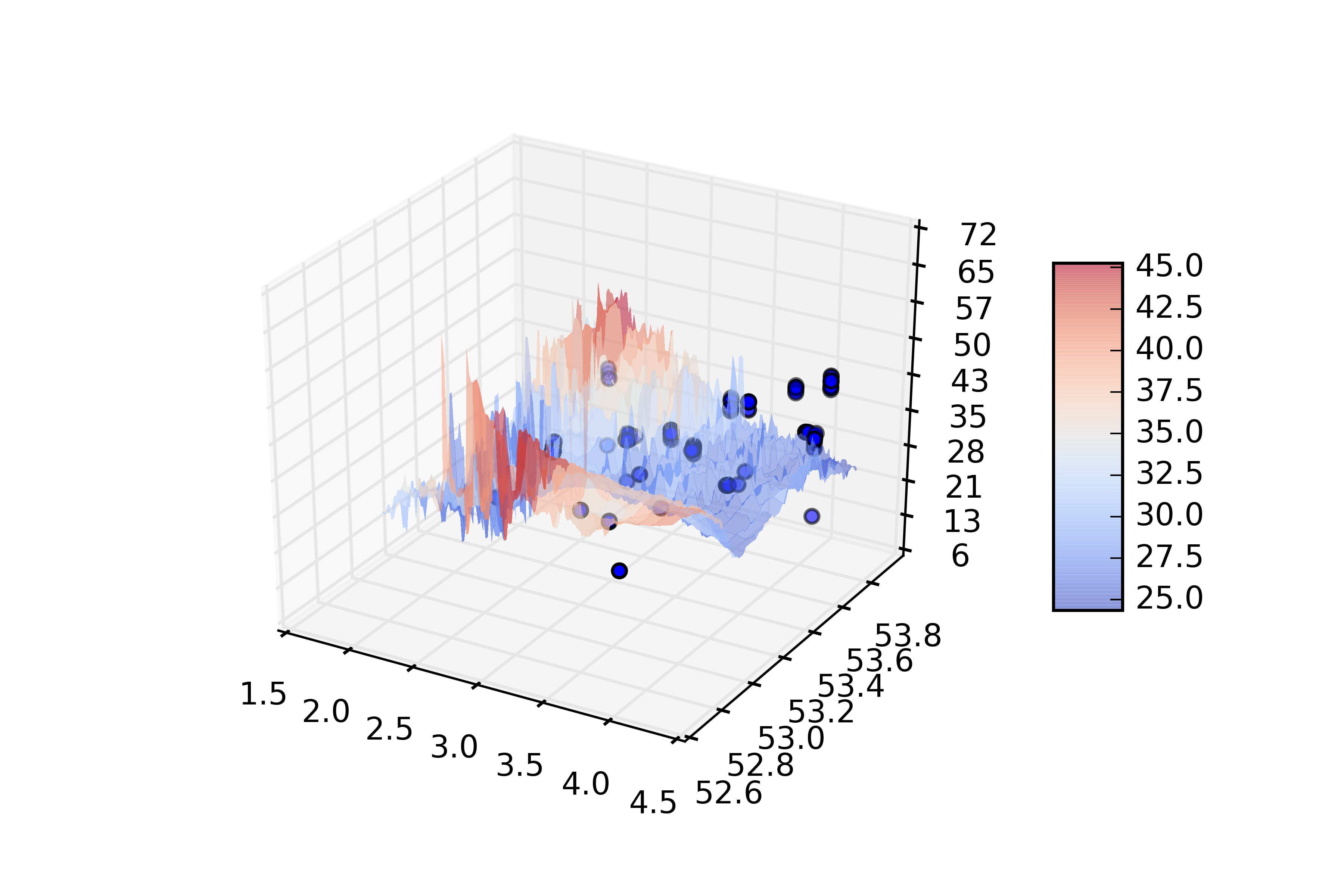
Code Block language py title Plotting the data img = theImage (see above) bbox = requestbbox (see above) def plotraster(img,df,resx,resy,bbox): from mpl_toolkits.mplot3d import Axes3D import matplotlib.pyplot as plt from matplotlib import cm from matplotlib.ticker import LinearLocator, FormatStrFormatter import numpy as np fig = plt.figure() ax = fig.gca(projection='3d') #plot the vectors xv = df['Longitude'] yv = df['Latitude'] zv = df['MaximumDepth'] ax.scatter(xv,yv,zv,label=df['ObservedIndividualCount']) # Make data. X = np.linspace(bbox[0],bbox[2], img.shape[1]) Y = np.linspace(bbox[1],bbox[3], img.shape[0]) X, Y = np.meshgrid(X, Y) #R = np.sqrt(X**2 + Y**2) #Z = np.sin(R) Z = img # Plot the surface. surf = ax.plot_surface(X, Y, Z, cmap=cm.coolwarm, linewidth=0, antialiased=False,alpha=0.5) # Customize the z axis. ax.set_zlim(Z.min().round(), Z.max().round()) ax.zaxis.set_major_locator(LinearLocator(10)) ax.zaxis.set_major_formatter(FormatStrFormatter('%.f')) # Add a color bar which maps values to colors. fig.colorbar(surf, shrink=0.5, aspect=5) plt.savefig(r'd:\temp\emodnet\test.png',dpi=600) plt.show()The result is visualised in the following graph
...