Contents
Acknowledgements
- This tutorial on visualising data from the OpenDap server is developed by the Data and Knowledge Management section of the Building with Nature programme as a Deliverable of Workpackage DM 1.1
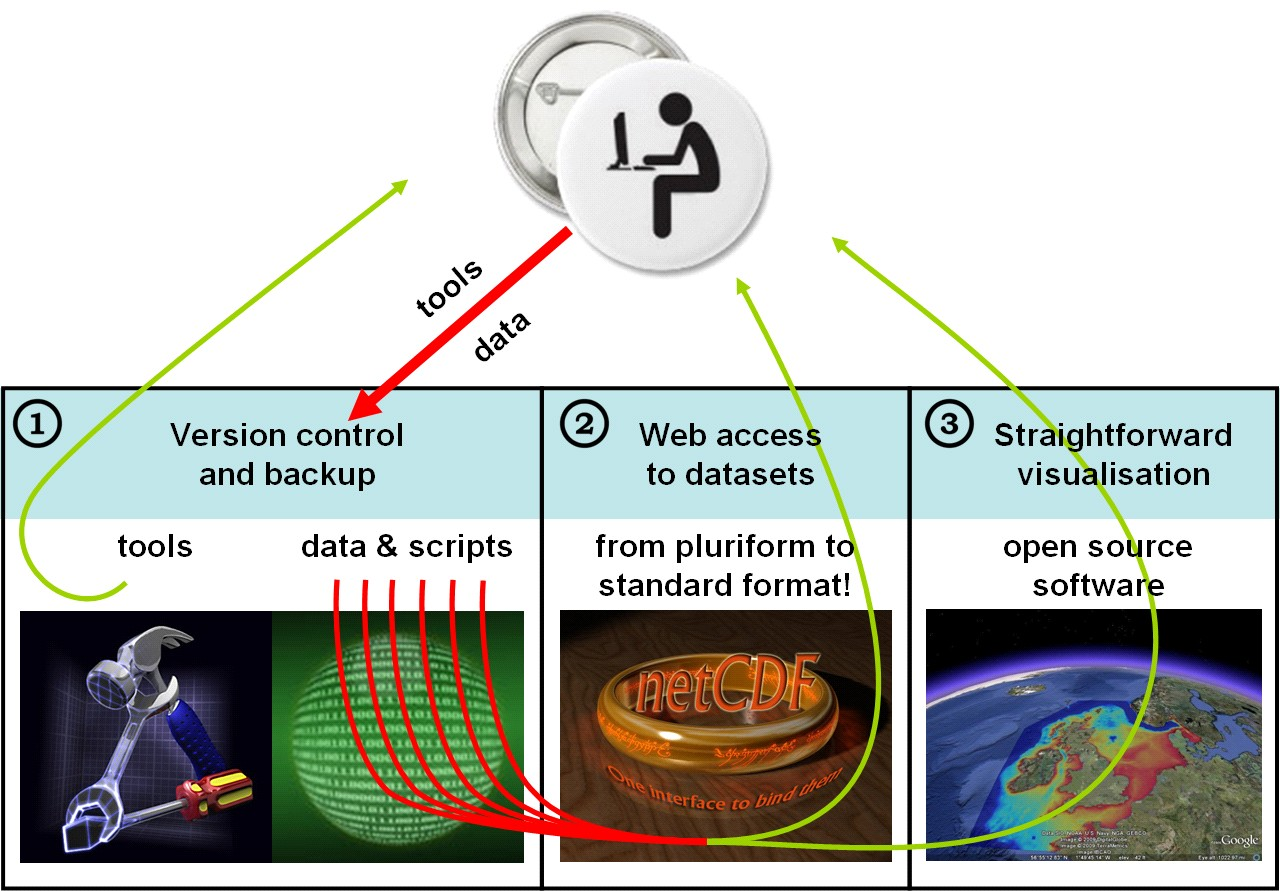
In this manual we describe how to use data from an OpenEarth system that consists of a Subverison repository, an OPeNDAP server and a server with Google Earth feeds.
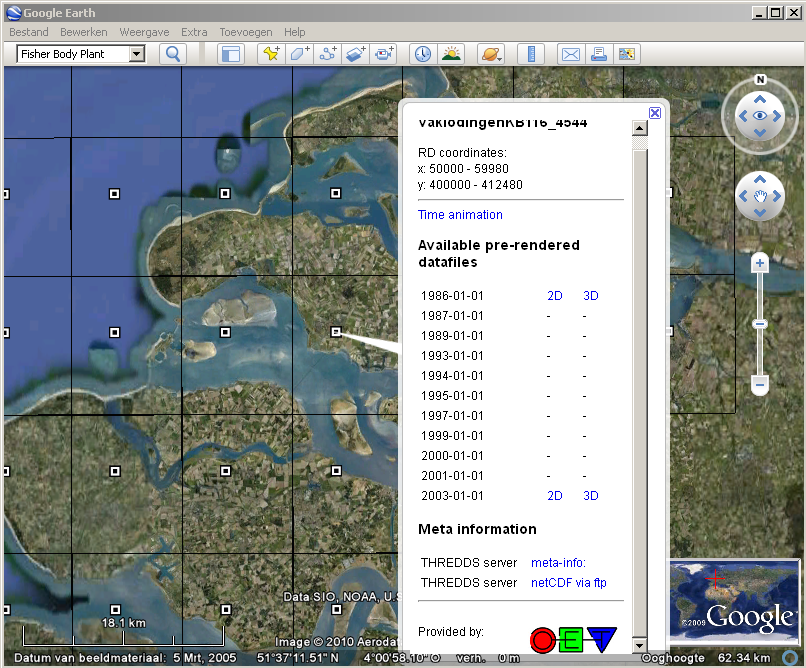
- Get the Google Earth file with the overview of the data from the kml server, e.g.:
- Maps
- Time series
- Rijkswaterstaat Waterbase (water levels, wave period/direction, water temperature, salinity, chlorofyll, suspended sediment)
- KNMI wind
- KNMI daily averaged meteo (wind, air temperature, sunshine, radiation, precipitation, air pressure, visibility, cloud cover, relative humidity, potential evapotransipration)
- Click on the placemark of the map or time serie you want to analyse, and then select the link THREDDS meta-info. Note: for the time series remember to drag the time ruler such that all times are available: drag the whole ruler first to the most recent data and then drag the left part of the ruler to the oldest date.
- In your browser the OPeNDAP meta-info page of the dataset will pop-up. You could also ave found this meta-info page by going down the OPeNDAP directory tree with a regular web brower staring at the top of the OPeNDAP three.
- Select all the in the box labeled Data URL.
- The 'DataURL' string can be used (copy, paste) to read the data directly from the web into various software packages, e.g.:
Examples for these packages are given below:
Accessing netCDF/OPeNDAP data with arcGIS
Buy arcGis 10 (9.2+) and use one of these options:
- use native arcGIS netCDF support (no curvilinear model grids!)
- http://webhelp.esri.com/arcgiSDEsktop/9.3/index.cfm?TopicName=Adding_netCDF_data_in_ArcGIS
- http://www.narccap.ucar.edu/data/gis-howto.html
- slides from ESRI at NASA HDF EOS workshop on improving arcGIS support for CF conventions http://hdfeos.org/workshops/ws15/agenda.php
- http://blogs.esri.com/esri/arcgis/2011/04/19/environmental-data-collector-extension-for-arcgis-10-0/
- download the free Environmental Data Connector EDC extension made by ASA on request of NOAA.
- MGET arcGis 10.1 addition: http://mgel.env.duke.edu/mget
Accessing netCDF/OPeNDAP data with ncBrowse
Download the free ncBrowse 1.6.3 from http://www.epic.noaa.gov/java/ncBrowse/. Note that the latest version is not the best to use. Use 1.6.3 if you only have classic netCDF3 files, use 1.6.5 only if you have netCDF3-64 bit offset files (neededf for files > 4 GB), but be aware that OPeNDAP functionality as explained in this tutorial malfunctions. For Windows 10, you might need to install the program with Compatibility mode set to Windows 7.
Right-click on the setup file and click on ‘Properties’.
Click on the ‘Compatibility’ tab and check the box ‘Run this program in compatibility mode for’ and select Windows 8/8.1 operating system or Windows 7 from the drop down menu and proceed with the installation.
For feed-back, please join the mailinglist ncbrowse@noaa.gov. The developer is aware of the plotting bugs in 1.6.6.
version | released | netCDF3 | netCDF3-64 bit offset | netCDF4 | OPeNDAP | plotting |
|---|---|---|---|---|---|---|
1.6.3 | 2006-aug-24 |
| ||||
1.6.5 | 2012-jun-15 | |||||
1.6.6 | 2013-apr-24 |
| ||||
1.6.7 | 2013-may-20 |
|
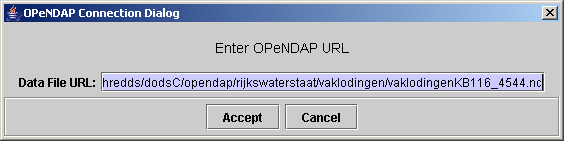
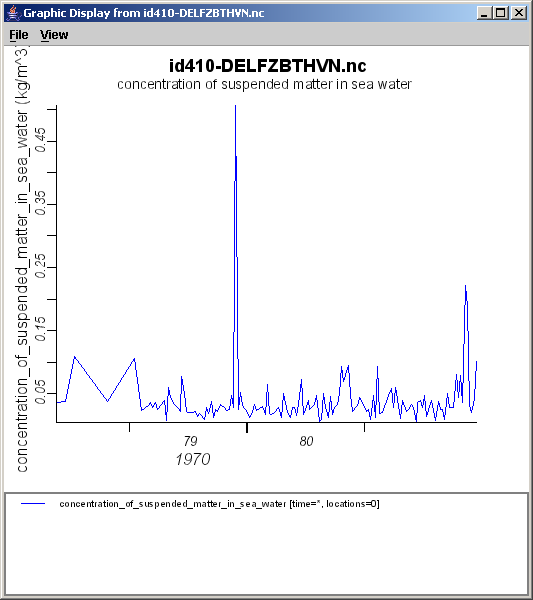
- select menu file > OPeNDAP
and paste the url you just copied. The viewer will now download the meta-information from the OPeNDAP server. Even if when the underlying file would be a hundreds of GB, this step is always very fast.
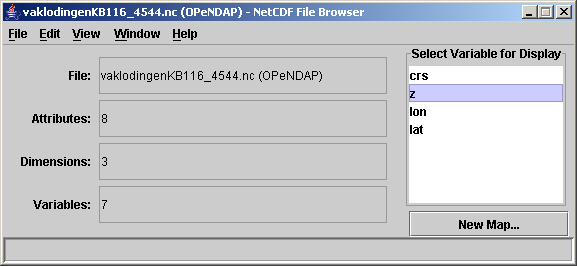
- select (double-click) the variable you want to plot, e.g. z.
- Define the data subset to get. Set the x dimension as x-axis, and y the y-dimension as y-axis. Not until you press graph variable the viewer will actually get the data. Even if when the underlying file would be a hundreds of GB, this step can be pretty fast provided the subset you chose is not too big. Only when you make too big a selection, ncBrowse will ill-behave. This will happen for instance with the 100 + years of tidal elevation dataset at station DelftZijl Buitenhaven. This selection step is the essential difference with the approach of downloading a netCDF file and then using ncBrowse to view that local file from your harddisk. With OPeNDAP you do not have to download an entire file before you can view it, because you can request only a specific slice, or a subset of a matrix.
- et voila (epxlore do the excellent zoom functionality of ncbrowse, especially the time-axis)
Accessing netCDF/OPeNDAP data with Matlab
Since Matlab R2012a native OPeNDAP support has been added to the Matlab netcdf package that exists since R2008b+. (For older Matlab releases, get snctools which is shiiped with OpenEarthTools) From OpenEarthTools run the following set-up function to add all relevant paths:
run('...\openearthtools\matlab\oetsettings.m')
- Go to an OPeNDAP server (e.g. http://opendap.deltares.nl) and pick a netCDF file by copying the contents of the Data URL box.
Define the associated url you just copied.
url_grid = 'http://opendap.deltares.nl/thredds/dodsC/opendap/rijkswaterstaat/vaklodingen/vaklodingenKB116_4544.nc' url_time = 'http://opendap.deltares.nl/thredds/dodsC/opendap/rijkswaterstaat/waterbase/concentration_of_suspended_matter_in_water/id410-DELFZBTHVN.nc'
and select the variable you want to plot by looking at the file contents
ncdisp(url_grid) ncdisp(url_time)
Define the data subset to get. Not until you issue the nc_varget you will actually get the data. Even if when the underlying file would be a hundreds of GB, this step can be pretty fast provided the subset you chose is not too big. Only when you make too big a selection, Matlab will take long. This will happen for instance with the 100 + years of tidal elevation dataset at station DelftZijl Buitenhaven. This selection step is the essential difference with the approach of downloading a netCDF file and then using Matlab to view that local file from your harddisk. With OPeNDAP you do not have to download an entire file before you can view it, because you can request only a specific slice, or a subset of a matrix.
G.x = ncread(url_grid,'x'); G.y = ncread(url_grid,'y'); G.z = ncread(url_grid,'z',[1 1 1],[Inf Inf 1]); % get only one timestep (here: 1st), but all x and y, note: ncread is zero based. note: matlab has different dimension order than other netCDF tools: t is last here
T.t = ncread_cf_time(url_time,'time'); % adapt reference date of 1970 in netCDF to Matlab reference date of 0000 T.eta = ncread(url_time,'concentration_of_suspended_matter_in_water');
plot ...
pcolorcorcen(G.x,G.y,G.z') axis equal axis tight tickmap('xy') grid on xlabel('x') ylabel('y') colorbarwithtitle('z')figure plot(T.t,T.eta) datetick('x') grid on- et voila
Download the code of this Matlab example (repos,OPeNDAP_access_with_Matlab_tutorial.m)
See also: Accessing netCDF/OPeNDAP data with python, Accessing netCDF/OPeNDAP data with R,
curvilinear model data subsetting with python
Accessing netCDF/OPeNDAP data with Python
Get PythonXY, add the netCDF4 package (PythonXY installer). Not all netCDF4 packages have been compiled with opendap yet. Therefore, manually also install the pydap package with easy_install.
python # start python in DOS shell to install easy_install >>> from ez_setup import use_setuptools >>> use_setuptools() >>> exit() # now easy_install works outside python easy_install Paste # sometimes Paste is problematic, so let's install itmanually easy_install Pydap
OpenEarthTools provides a module opendap.py that makes pydap quack like netCDF4 (repos, manual download) so you can talk directly to opendap data via the web. Now execute the following Python lines, or download the full example code (repos,manual download):
#!/usr/bin/env python # Read data from an opendap server import netCDF4, pydap, urllib import pylab, matplotlib import numpy as np from opendap import opendap # OpenEarthTools module, see above that makes pypdap quack like netCDF4
- Go to an OPeNDAP server (e.g. http://opendap.deltares.nl) and pick a netCDF file by copying the contents of the Data URL box.
- Define the associated url you just copied.
url_grid = r'http://opendap.deltares.nl/thredds/dodsC/opendap/rijkswaterstaat/vaklodingen/vaklodingenKB116_4544.nc' url_time = r'http://opendap.deltares.nl/thredds/dodsC/opendap/rijkswaterstaat/waterbase/concentration_of_suspended_matter_in_water/id410-DELFZBTHVN.nc'
- Extract the data.
# Get grid data grid = opendap(url_grid) G_x = grid.variables['x'] G_y = grid.variables['y'] G_z = grid.variables['z'] G = {} # dictionary ~ Matlab struct G['x'] = G_x[:].squeeze() G['y'] = G_y[:].squeeze() G['z'] = G_z[1,:,:].squeeze() # download only one temporal slice # represent fillValue from data as Masked Array # the next release of netcdf4 will return a masked array by default, handling NaNs # correctly too (http://code.google.com/p/netcdf4-python/issues/detail?id=168) G['z'] = np.ma.masked_invalid(G['z'])# Get time series data time = opendap(url_time) T_t = time.variables['time'] T_z = time.variables['concentration_of_suspended_matter_in_water'] T = {} # dictionary ~ Matlab struct T['t'] = netCDF4.num2date(T_t[:], units=T_t.units) T['z'] = T_z[:].squeeze() - plot ...
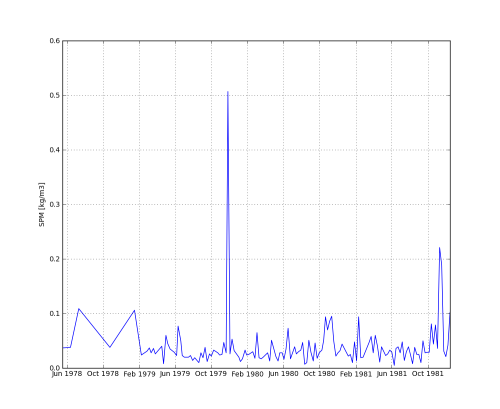
# plot grid data matplotlib.pyplot.pcolormesh(G['x'],G['y'],G['z']) pylab.xlabel('x [m]') pylab.ylabel('y [m]') matplotlib.pyplot.colorbar() matplotlib.pyplot.axis('tight') matplotlib.pyplot.axis('equal') matplotlib.pyplot.title(url_grid) pylab.savefig('vaklodingenKB116_4544') # plot time series data pylab.clf() matplotlib.pyplot.plot_date(T['t'], T['z'], fmt='b-', xdate=True, ydate=False) pylab.ylabel('SPM [kg/m3]') matplotlib.pyplot.title(url_time) pylab.savefig('DELFZBTHVN') - et voila
Download the code of this python example (repos,manual download) + opendap.py pydap2netCDF4 module (repos, manual download).
See also: OPeNDAP subsetting with python,Accessing netCDF/OPeNDAP unstructured grids with python, Accessing netCDF/OPeNDAP data with R, Accessing netCDF/OPeNDAP data with Matlab, curvilinear model data subsetting with python
Accessing netCDF/OPeNDAP data with R
Get R, which includes several netCDF4 packages.
require(ncdf)
- Go to an OPeNDAP server (e.g. http://opendap.deltares.nl) and pick a netCDF file by copying the contents of the Data URL box. Because the netcdf packages for windows are not yet opendap-enabled, download them.
- Define the associated url you just copied.
A complete linux image with the R netcdf package compiled with OPeNDAP is available upon request from ""adaguc "at" knmi.nl"".
url_grid <- "http://opendap.deltares.nl/thredds/fileServer/opendap/rijkswaterstaat/vaklodingen/vaklodingenKB116_4544.nc" # note: netcdf4 does not work on windows R url_time <- "http://opendap.deltares.nl/thredds/fileServer/opendap/rijkswaterstaat/waterbase/concentration_of_suspended_matter_in_sea_water/id410-DELFZBTHVN.nc" download.file(url_grid, "vaklodingenKB116_4544.nc", method = "auto", quiet = FALSE, mode="wb", cacheOK = TRUE) download.file(url_time, "id410-DELFZBTHVN.nc", method = "auto", quiet = FALSE, mode="wb", cacheOK = TRUE)
- Extract the data.
grid.nc <- open.ncdf("vaklodingenKB116_4544.nc") # look what's in there... grid.nc # Get grid data G.x <- get.var.ncdf(grid.nc,'x') G.y <- get.var.ncdf(grid.nc,'y') # get only first timestep G.z <- get.var.ncdf(grid.nc,'z')[,,1] # to get a black background, and set the scale of depth values to start from 0. G.z[G.z == -9999] <- 0 # image.plot needs sorted x- and y-values; # as y-values are descending, the order is reversed here... G.y <- rev(G.y) G.z <- G.z[,length(G.y):1]time.nc <- open.ncdf("id410-DELFZBTHVN.nc") # look what's in there... time.nc T.t <- get.var.ncdf(time.nc,'time') T.eta <- get.var.ncdf(time.nc,'concentration_of_suspended_matter_in_sea_water') - plot ...
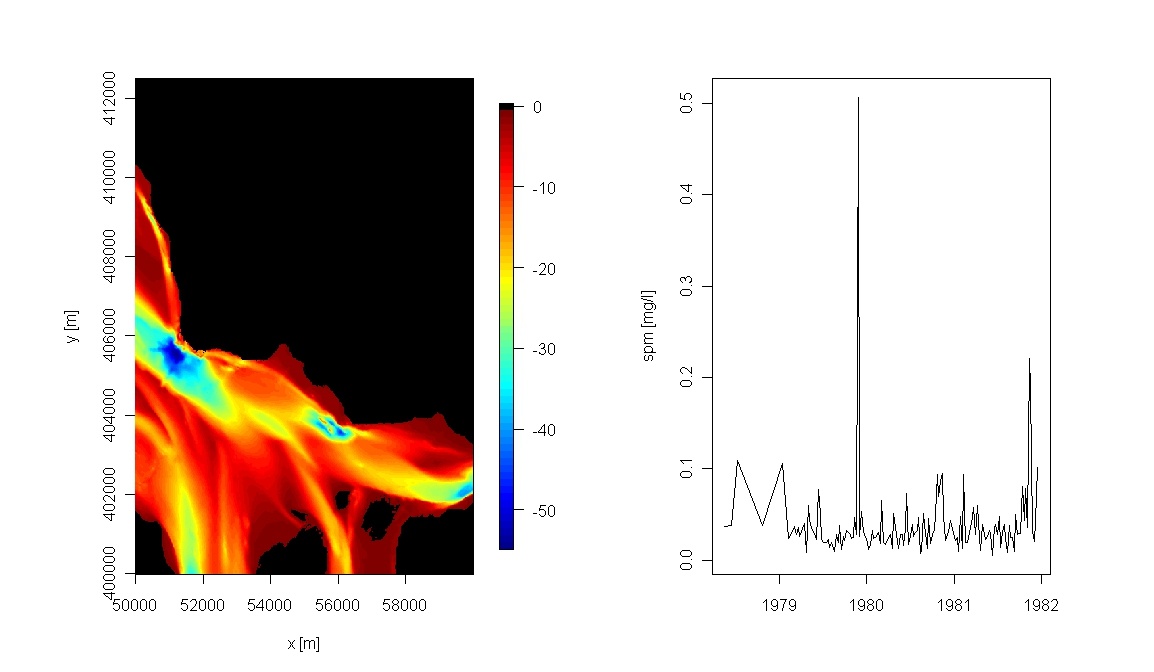
# R-package fields provides nice image facilities and color schemes par (mfrow = c(1,2)) library(fields) image.plot(G.x,G.y,as.matrix(G.z), col = c(tim.colors(),"black"), xlab = "x [m]", ylab = "y [m]")plot(as.Date(T.t, origin="1970-01-01"), T.eta, type = "l", ylab = "spm [mg/l]")
- et voila
Download the code of this R example (repos,manual download), which was provided by Karline Soetaert and Tom van Engeland.
See also: Accessing netCDF/OPeNDAP data with python, Accessing netCDF/OPeNDAP data with Matlab, PostgreSQL access with R, OPeNDAP subsetting with R
Accessing netCDF/OPeNDAP data with Delft3D-Quickplot.
Get Delft3D (code is available as open source since jan 1st 2011). (Download Delft3D-Quickplot-OPeNDAP manual.)
- select menu file > url and paste the url you just copied. Delft3D-Quikcplot will now download the meta-information from the OPeNDAP server. Even if when the underlying file would be a hundreds of GB, this step is always very fast.
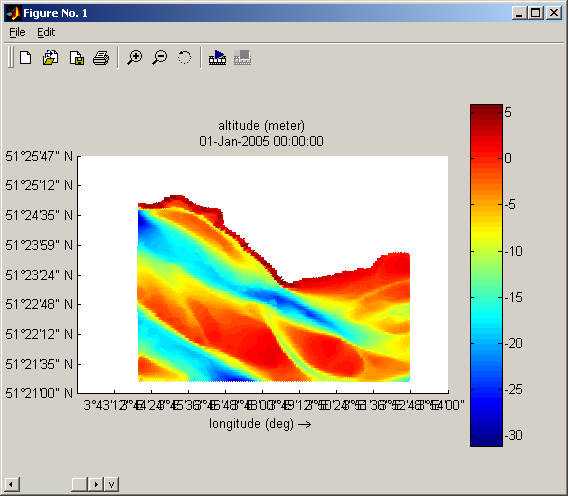
- Select the variable you want to plot by looking at the file contents
- Define the data subset to get.
- et voila